No-code deep learning with Jiva.ai
A foray into no-code AI and my thoughts on its utility for scientists.
Dr Chetan Kaher is on a mission to empower every clinician to build AI models. We first met at an NHS event back in 2019. By day, he led a successful dental practice in London. By night, he and co-founder Dr Manish Patel were building Jiva.ai, a platform that lets anyone build artificial intelligence (AI) without writing code. “It’s easy to forget that once upon a time, you needed a computer science degree to build a website”, he told me, “But today, WordPress and Wix let you point and click your way to a beautiful-looking webpage. Why can’t we do the same artificial intelligence”? AI sounded like science fiction at that time (the Dark Ages, before ChatGPT), and pointing and clicking your way to AI sounded wishful, so I wished him well.
We crossed paths again early in 2023. He failed to mention that Jiva.ai now boasts alumni status at accelerator programmes like KQ Labs and P4 Precision Medicine, a long list of partner institutes and a successful round of seed funding. I learned all this later. Instead, he shared that Jiva.ai was being used beyond the clinic. Working in the life sciences, the need for tools like this is obvious, where groundbreaking AI like AlphaFold have been birthed from the rising tide of biological data and where a digital skills gap threatens to bottleneck future discoveries. I wanted to know if their no-code AI platform would be good for scientists.
A blood test for smokers
We can expect a tool like Jiva.ai to let non-coders replicate analyses. In my field of cancer genomics, we know that smoking can lead to lung cancer because of the specific ways it damages DNA. Scientists can measure features like DNA methylation from our blood and build models that can tell how much we smoke, how often we smoke, and predict our risk of developing cancer [1]. A recent study by Vidaki., A. et al. (2023) reported 13 DNA marks that could predict smoking status with 78% accuracy across 232 patients [2].
For comparison, I first reproduced the paper’s results with code using the data they made public. With a virtual computer from paperspace.com, I installed the Python Pytorch library and ran a feed-forward neural network (notebook available here). With a good knowledge of Python and the Pytorch library, I estimate most people could get this working in a few hours. After training on a subset of data, my model had a 72.86% accuracy on the holdout test set. Now, can we get there with Jiva?
No-code smoking prediction with Jiva.ai
Jiva.ai runs as a simple web application. I opened the site, followed all the obvious signals to create a project, and uploaded the DNA methylation measurements from the paper by Vidaki., et al. (2023). Jiva.ai showed me the distribution of each variable in the dataset. From this view, we could immediately see the variable for our AI model to predict (Smoking Status) and our DNA methylation features, each represented by a ‘cg’ identifier.
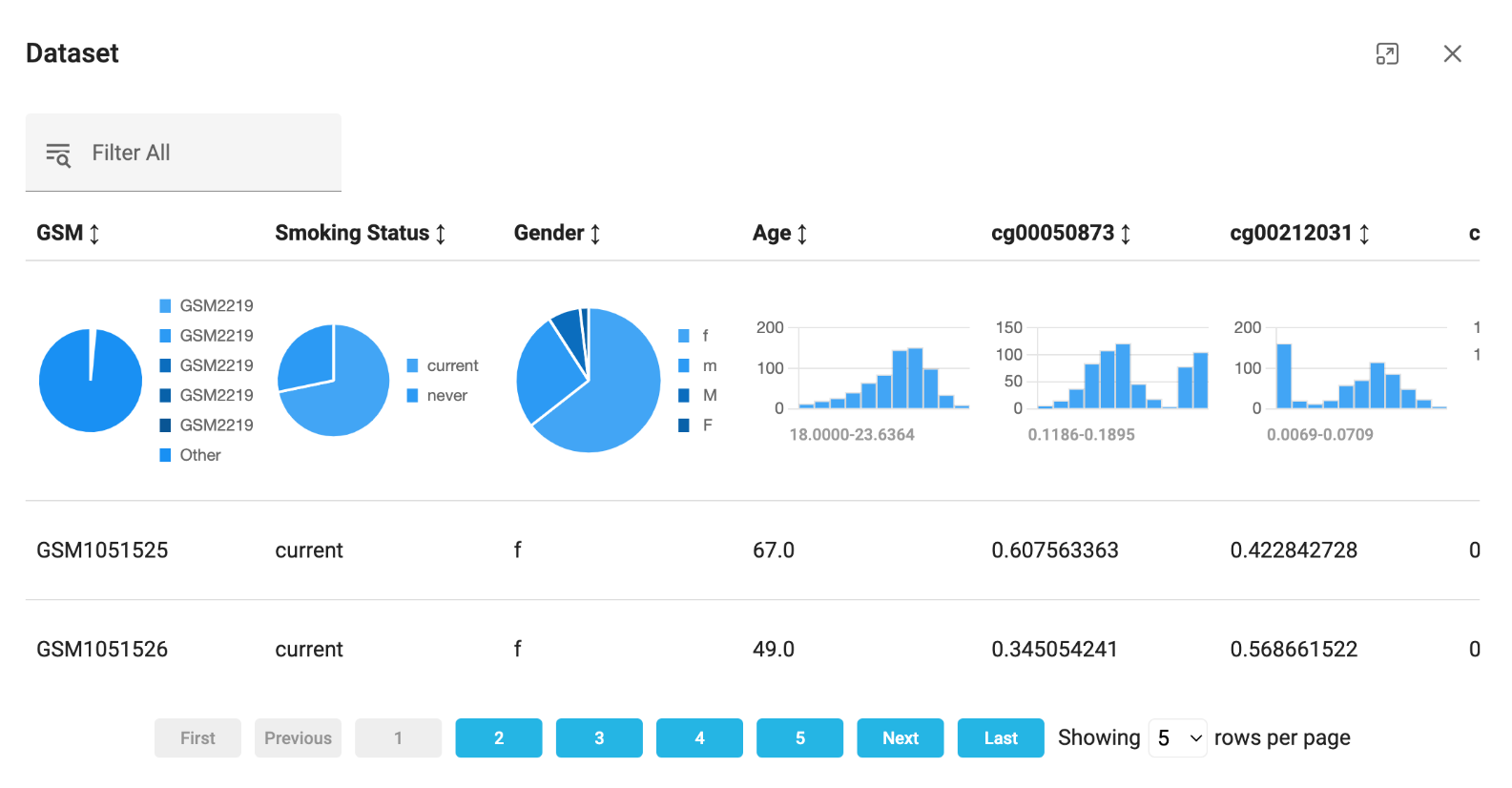
Once data is up on Jiva.ai, you can start building a workflow. It gave me a canvas to place and connect nodes together. The result was a visual representation of the deep learning model. Interacting with the platform is like putting together a puzzle or moving a deck of cards around a table.
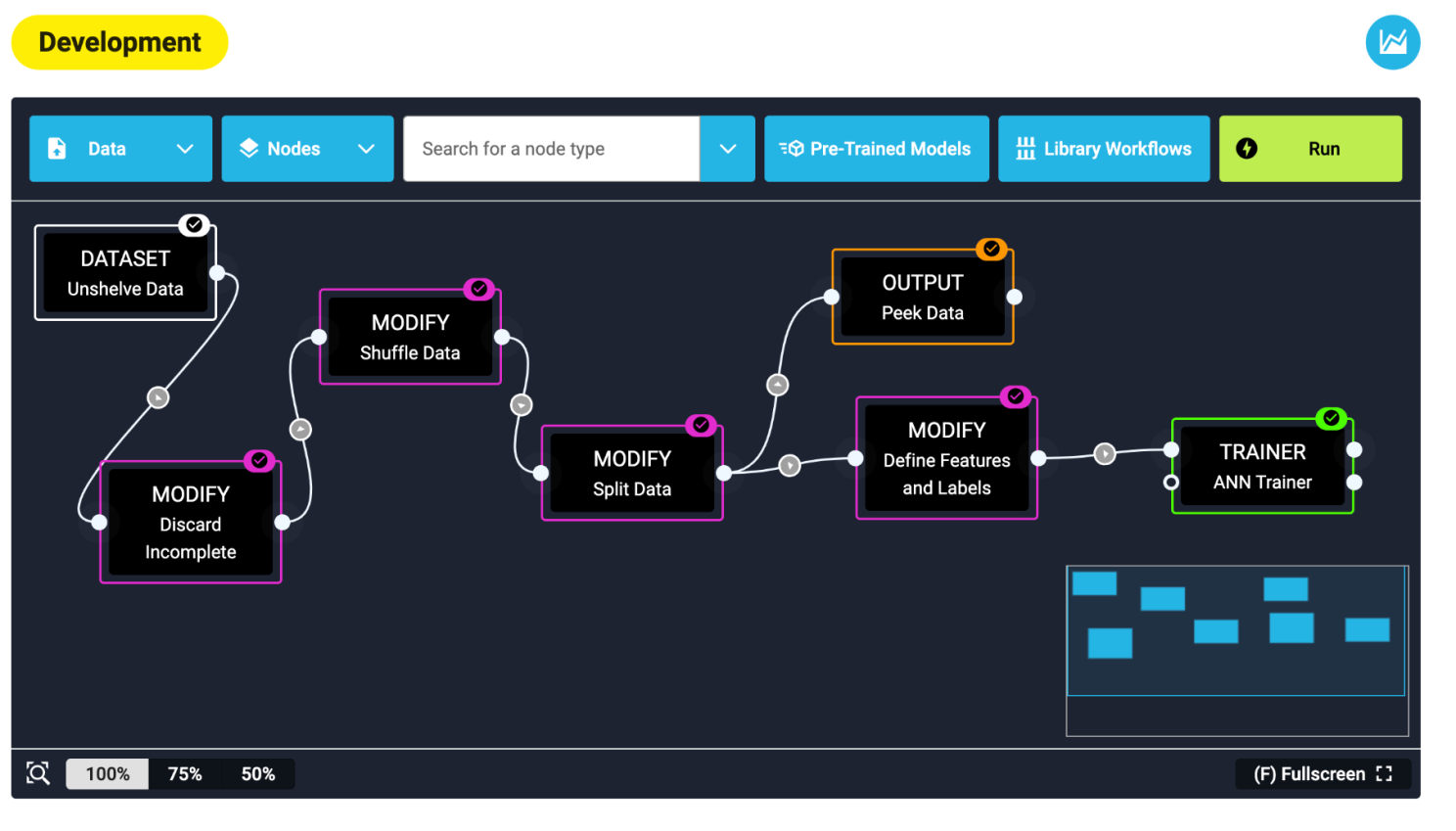
Nodes are easy enough to find, grouped sensibly by what they do: data processing, scaling, feature selection and model training. I piped together a shuffler, data splitter, feature selector and finally, an artificial neural network trainer. I hit the bright green ‘Run’ button and…. Error! My pipeline failed. Jiva.ai returns a log file with every training run, which I consulted to learn that I needed to add a node to remove empty records from my data. After doing so, I hit run again, and voila! A green tick. I wouldn’t blink twice if I saw an error while coding, but this was a surprise from a no-code platform, like getting an error from your word processor. However, at its heart, Jiva.ai is an interface to Python code which remains forever out of sight.
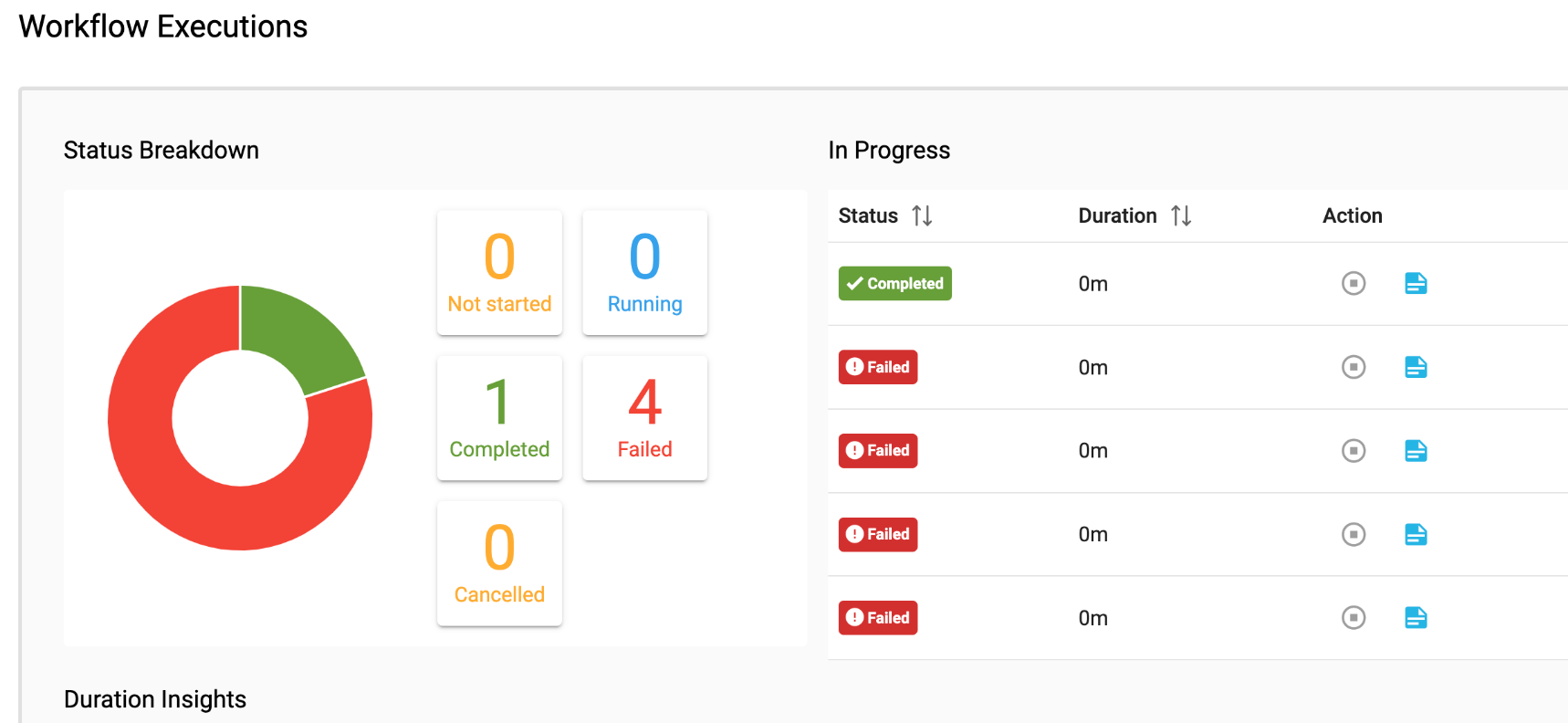
Altogether, this took me around half the time as with Pytorch. Most of the time was spent searching through Jiva’s menus for the correct component name, whereas when I was coding, my time was spent double-checking the Pytorch documentation. As a quick-and-easy prototype, the model built with Jiva achieved 69.5% accuracy on the test dataset without any significant tweaks to the default neural network trainer and without writing a line of code.
What do no-code AI platforms mean for scientists?
Enabling everyone to train and deploy AI models is a profound concept. No-code AI fills a skills gap where the global demand for deep learning expertise outpaces our ability to meet it. The striking feature of Jiva.ai is how quickly I could get going without touching anything resembling a user manual. Every coding framework for deep learning demands that you read documentation, and if it’s sufficiently complex, this can feel like learning a new language. With no-code platforms, you’re running straight out of the gate.
But is this useful for scientists? Scientists are looking to build AI models using their experimental data. In this example, that means measuring DNA marks from blood samples and using them to predict smoking status. The next step is understanding why. What is it about smoking that targets these specific parts of DNA? Explainability is a big challenge in AI. Any no-code platform that gives users these tools would set itself apart from the competition as far as scientists are concerned.
Lowering the barrier to building models might lead to more scientific discoveries. The platform’s price point will determine how low that barrier is. There’s an exciting equity angle if platforms like Jiva.ai too if they can bring AI tools to parts of the world that can’t afford AI developers. In the long run, this could be a gateway to learning how to code. I realised this when using Jiva.ai, as node names like ‘one-hot encoder’ and ‘artificial neural network trainer’ implicitly need users to understand the nuts and bolts of building models. At the same time, no-code AI could be a rapid prototyping tool for AI-savvy scientists looking to build models.
As scientists, we never trust a result that we can’t reproduce. Sharing models is the backbone of reproducing and using AI research. It should be just as easy to deploy or use an existing model as it is to build it. Hugging Face built a $2B platform for providing this service and is already a standard for hosting deep learning models with code, so no code platforms for science should take note.
Viva la Jiva
Speaking to Chetan and Manish, I get the impression that Jiva.ai has been a labour of love. The world moves fast. But they find themselves in precisely the right place in time. I went from sceptical, to curious, to hopeful. Jiva.ai has a long road ahead, but their work so far is a sign of things to come. Your ability to build AI models will soon be a few clicks away. Just ask your dentist.
Acknowledgements
Thanks to Mohammed Shaaban for providing feedback on early drafts.Disclaimer: Disclaimer: I have no professional affiliation or financial relationship with Jiva.ai. This article was written independently and sent to the co-founders as a courtesy. All views are my own.
References
-
Yousefi PD, Suderman M, Langdon R, Whitehurst O, Davey Smith G, Relton CL. DNA methylation-based predictors of health: applications and statistical considerations. Nat Rev Genet. 2022;23: 369–383. doi:10.1038/s41576-022-00465-w
-
Vidaki A, Planterose Jiménez B, Poggiali B, Kalamara V, van der Gaag KJ, Maas SCE, et al. Targeted DNA methylation analysis and prediction of smoking habits in blood based on massively parallel sequencing. Forensic Sci Int Genet. 2023;65: 102878. doi:10.1016/j.fsigen.2023.102878


